Method Development for Microbial Ecology
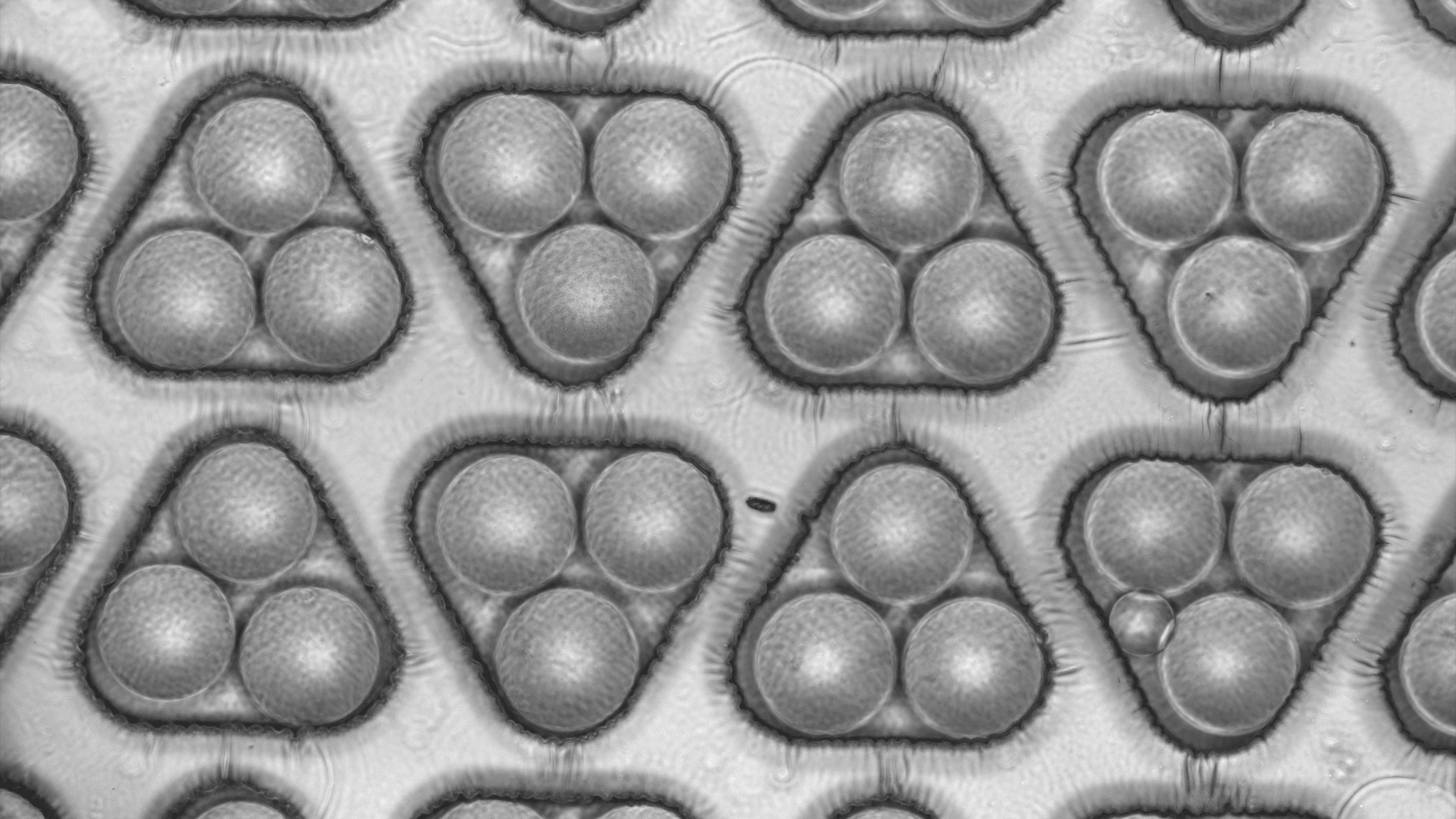
Close-up view of a kChip filled with droplets in which growth of bacterial cells can be monitored at high-throughtput.
Developing and implementing new methods enables us to overcome existing limitations and obtain more accurate, comprehensive data. We are mostly working on methods, tools and software related to microfluidics and live-cell imaging, inluding:
- MIDAP, a pipeline for automated segmentation and tracking of bacterial cells in time-lapse microscopy image series; a selection of state-of-the-art segmentation and tracking methods are provided from which the user can select the most suited ones for their data. Available on external page GitHub (with Lukas von Ziegler from ETH SIS)
- Microfluidic designs to study microbial interactions (with Stefano Ugolini)
- A combined microfluidics and nanoSIMS method to couple analyses of single-cell growth and metabolic activity (Benjamin with Glen D'Souza, Fabai Wu and the group of Victoria Orphan)
- A device to perform anaerobic microfluidics-based live-cell imaging (with Alyson Hockenberry, Markus Arnoldini and Susan Schlegel)
- The kChip, a droplet-based platform designed for the rapid, massively parallel construction and screening of synthetic microbial communities (Anna, originally developed by the external page Blainey lab)